PCR Arrays for Quantitative PCR
Measuring gene expression by PCR array analysis is an excellent way to gain insight into differentially expressed genes associated with disease and other phenotypes

Measuring gene expression by PCR array analysis is an excellent way to gain insight into differentially expressed genes associated with disease and other phenotypes
PCR arrays provide a simple and powerful method to analyze gene expression for hundreds of genes simultaneously. These assays require access to a 96 or 384-well real time PCR instrument and a plate of primer sets designed to amplify specific cDNAs representing a specified pathway or group of genes/biomarkers. Specialized PCR arrays also enable the measurement of miRNA, circRNA, DNA copy number and mutation status. Detection may be done using intercalation dyes (ie. SYBR Green) or hybridization/hydrolysis probe chemistries.
The advent of high throughput gene expression techniques such as microarray-based transcriptional profiling and RNA-Seq provide a robust method for identifying differentially expressed genes in a cell or RNA sample. Though these techniques are expensive and have quantitative limitations including inaccuracies in measuring fold differences due to signal suppression (microarray) and variability in results due to library preparation (RNA-Seq), one can obtain a large list of potential genes that require further validation. Validation using PCR arrays are ideal for this step as one can perform quantitative gene expression analysis on a focused set of genes for multiple replicates and multiple samples at a relatively low cost. Quantitative real-time PCR is considered a more accurate, reliable, and reproducible method for this analysis.
Methodology
A typical PCR array reaction may be done with as little as 1-10 ng of RNA (cDNA) per plate. Careful RNA isolation and cDNA preparation is required to obtain the best results. RNA isolation should yield high quality RNA as assessed by the intactness of the 18S and 28S RNA bands on an agarose gel. Treatment of the purified RNA with DNase I will also guarantee that no contaminating genomic DNA is present in the assay. Reverse transcription may be performed with oligo-dT or random primers and the resulting cDNA should be stored at -80C for medium to long-term as it tends to be labile at warmer temperatures.
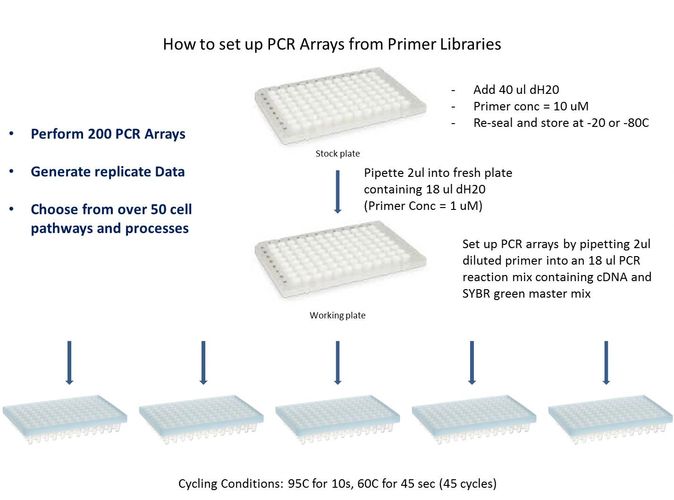
Setting up PCR arrays is simple, especially if one has access to multichannel pipettes. Prepare core solutions when possible to minimize well-to-well variability in the assay. First, prepare a core solution containing water, qPCR Master Mix and cDNA, enough to fill all wells of the plate plus 10% extra to accommodate any pipetting errors. Once this solution is evenly distributed on the plate, one can pipette the primer sets into each well (unless they are already present in a pre-aliquoted plate). Simply seal the reaction plate with an appropriate qPCR optically transparent tape sheet and place in the qPCR instrument. If preparing multiple reaction plates at the same time, seal and store the plates at -80C until they can be run. See the figure above for setting up PCR arrays from a PCR primer library.
We use cookies to analyze website traffic and optimize your website experience. By accepting our use of cookies, your data will be aggregated with all other user data.